Juan Viguera Diez
@viguera10
PhD student at AstraZeneca and Chalmers University of Technology. Machine Learning for natural sciences (Drug discovery). @viguera10.bsky.social
ID: 1547191021800493056
13-07-2022 12:08:44
74 Tweet
110 Followers
181 Following

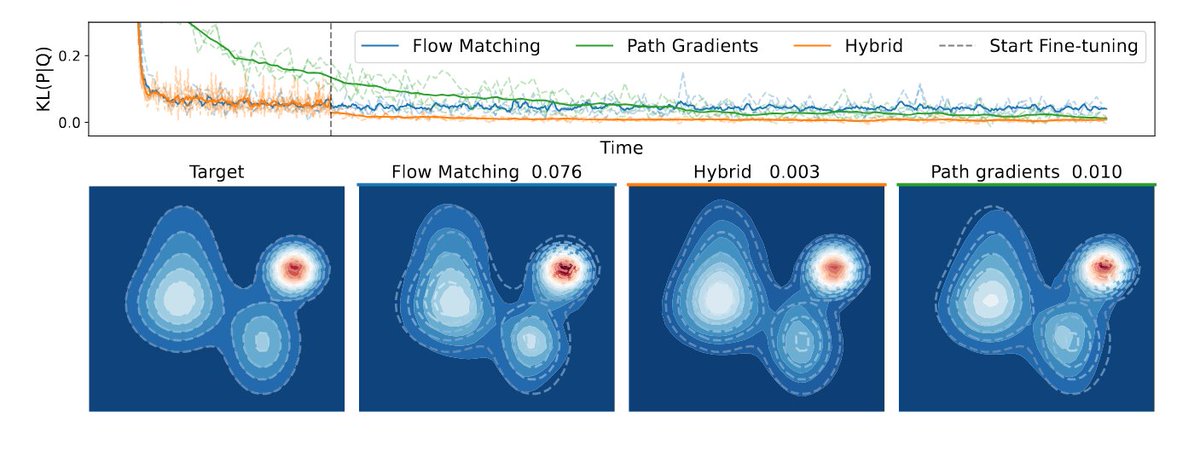
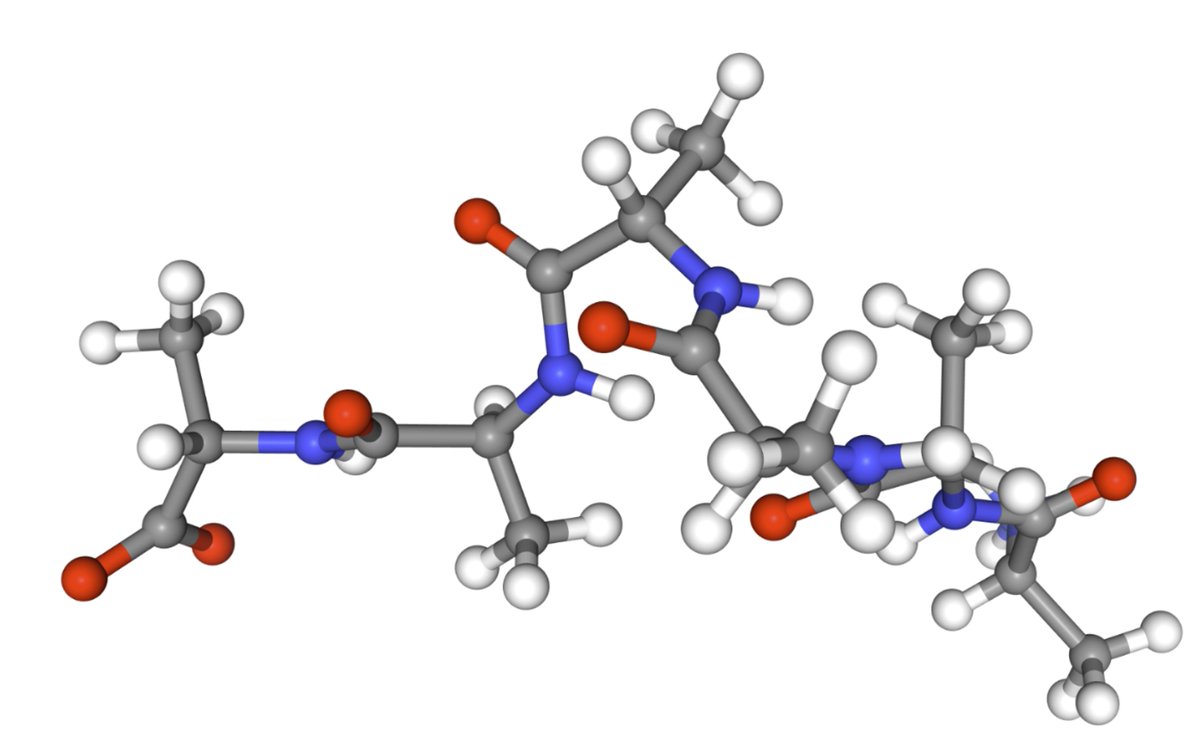
New preprint! 🚨 We scale equilibrium sampling to hexapeptide (in cartesian coordinates!) with Sequential Boltzmann generators! 📈 🤯 Work with Joey Bose, Chen Lin, Leon Klein, Michael Bronstein and Alex Tong Thread 🧵 1/11




Registration for this years CHAIR Structured Learning Workshop is open. Speakers include: Klaus Robert Müller, Jens Sjölund, Alex Tong, Jan Stühmer Arnaud Doucet, Marco Cuturi, Marta Betcke, Elena Agliari, Beatriz Seoane, Alessandro Ingrosso, ui.ungpd.com/Events/60bfc7b…







Excited to release FORT, a new regression-based approach for training normalizing flows 🔥! 🔗 Paper available here: arxiv.org/abs/2506.01158 New paper w/ Oscar Davis Jiarui Lu Jian Tang Michael Bronstein Yoshua Bengio Alex Tong Joey Bose 🧵1/6






BioEmu now published in Science Magazine !! What is BioEmu? Check out this video: youtu.be/LStKhWcL0VE?si…

📢📢 "La-Proteina: Atomistic Protein Generation via Partially Latent Flow Matching" Fully atomistic. Partially latent. Structurally precise. Entirely generative. w/ Tomas Geffner*, Kieran Didi @ICLR*, et al. 📜 Project page & paper: research.nvidia.com/labs/genair/la… 🧵 Thread below... (1/n)

🚀 Come check our poster at ICML GenBio Workshop @ ICML25! We show that pretrained MLIPs can accelerate training of Boltzmann emulators — by aligning their internal representations. Coauthors PINEDE Lucas, Juno Nam, RGB Lab @ MIT (1/n)


Our development of machine-learned transferable coarse-grained models in now on Nat Chem! doi.org/10.1038/s41557… I am so proud of my group for this work! Particularly first authors Nick Charron, Klara Bonneau, Aldo S. Pasos-Trejo, Andrea Guljas.